Towards Systematic Models of Cellular Regulation with Systems Biochemistry
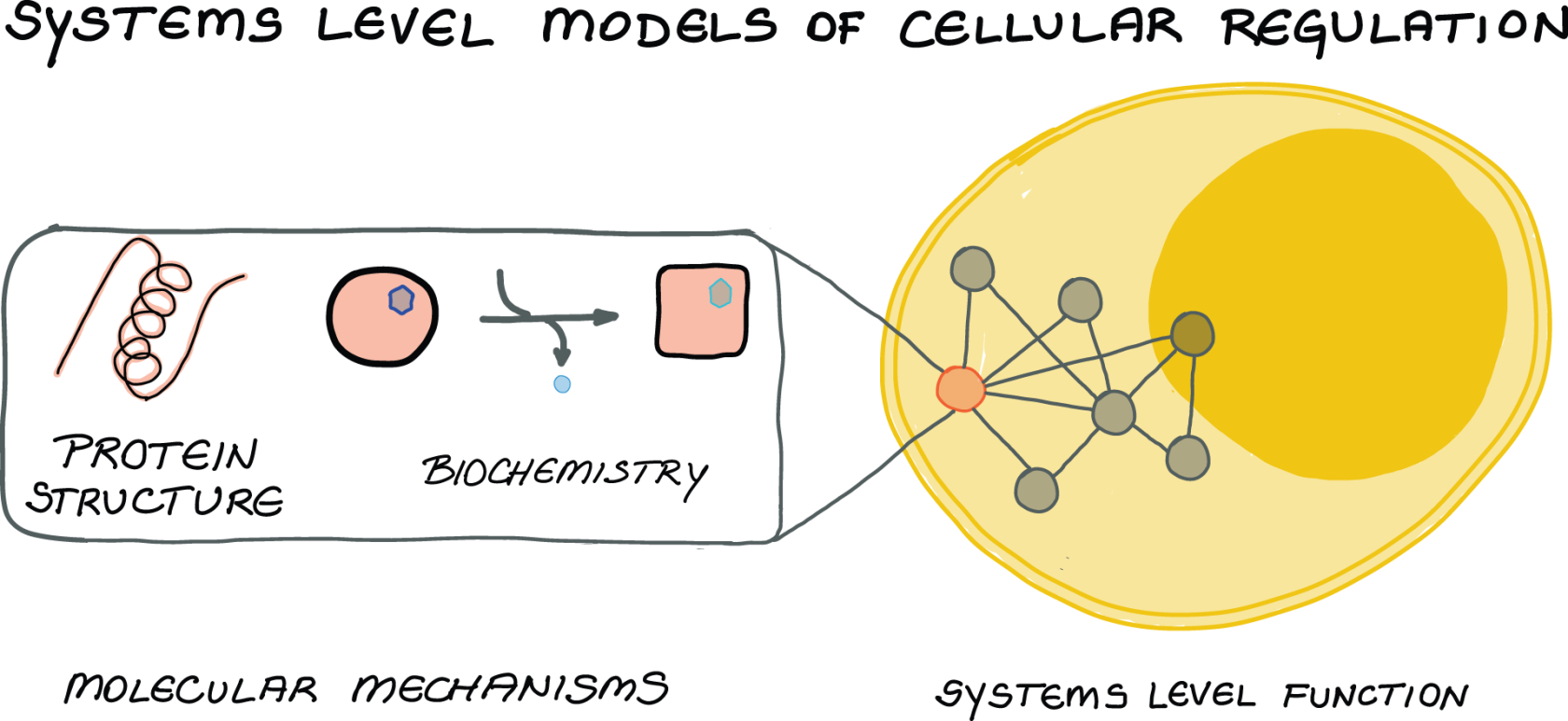
Our work combines “small-scale biology” of protein molecular mechanisms and “large-scale” network biology. Specifically, we seek to answer how structural and biochemical properties of signalling component, such as GTPases and protein kinases, encode their complex functions. How do the emergent properties of signalling proteins enable them to simultaneously, yet precisely, regulate different cellular processes?
We believe that a systems-view of protein function is essential for achieving truly predictive understanding of how mutations or therapeutic molecules affect the phenotype. This level of genotype-to-phenotype mapping could lead to better evaluation of disease mutations and effectiveness of drugs, but it requires an integrated approach of study that combines quantitative measurements of systems level cellular phenotype with fine-grained biophysical models of molecular mechanisms.
We use structural bioinformatics and evolutionary analysis to design targeted perturbations in signalling and regulatory proteins, and biochemically analyse their effects on the molecular function. Our lab also develops experimental and computational functional genomics methods that use these biochemically defined targeted perturbations to map functional interactions between and across signalling and regulatory pathways in cells.