Structural Studies of Eukaryotic Cells and Organelles
Our group uses structural approaches to address major questions in cellular structural biology including cytoskeletal organization and structure-based functional organization of the nuclear periphery. We combine several microscopic techniques such as high-end fluorescent microscopy, focused-ion beam milling, cryo-electron tomography (CET) and, most recently, atomic force microscopy (AFM), to study cellular processes. Additionally, we develope novel biochemical and structural tools to modify specific assemblies and allow a direct imaging of macromolecular complexes and their structural analysis in situ. In recent years we focus on the following projects:
Lamins are nuclear intermediate filaments (IFs) of metazoan cells
They assemble into fibrous structures that are positioned between the inner nuclear membrane and the peripheral chromatin, although a small fraction of lamins is present in the nucleoplasm. Lamins are required to maintain nuclear structure and, together with many interaction partners, are involved in most nuclear activities. Mutations in lamins cause a group of >14 distinct diseases called laminopathies. It is not clear how lamins are organized in vivo and how lamins’ mutations affect lamins’ functions. Results on the 3D organization of the Caenorhabditis elegans lamin provided the first structural fundament to lamin filament assemblied ex-vivo. Recently, we have provided the first view into lamin filaments in somatic and mammalian cells in situ and the first cellular alterations caused by a laminopathic mutation. The use of FIB-milling approach in conjunction with cryo-ET allows us to gain insight into the lamin organization in patient cells. Moreover we detected differences in the nuclear envelope organization as a function of releasing the mechanical pressure on the nuclear envelope by cell permeabilization and washing components of the cytoplasm.
In combination with mechanical studies, these results are a major step towards the understanding of lamin modus operandi and the special properties of nuclear lamin filaments.
Development of labeling approach and filaments structural determination for in situ structural analysis by CET
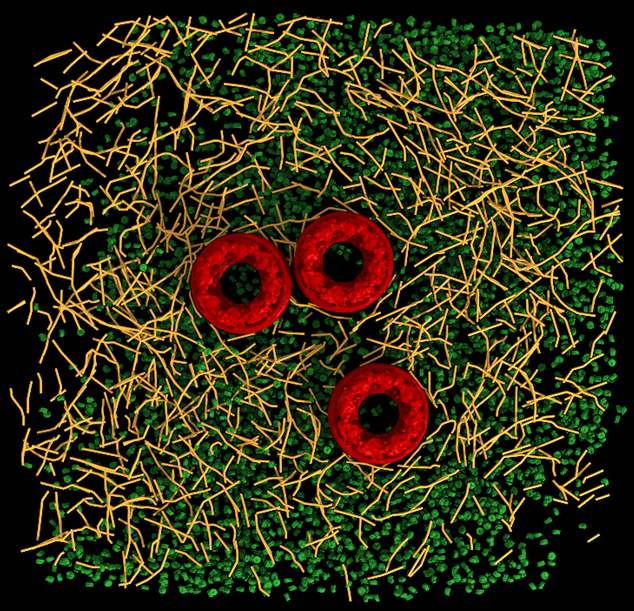
Recently, we synthesized a 2.2nm nanoparticles that can readily be conjugated to biological molecules, such as peptides and proteins, and be used to reveal the position of specific proteins in CRT with nanometric resolution. This allowed to identiy receptors in the plasma membrane and other protein, e.g. vinculin bound to actin. Moreover, we developed the means to determine the polarity of each actin filament in CET data. The protocol allows to map the directionality of the cytoskeleton from un-labeled cells. The actin filaments are reconstructed to sub-nanometer resolution which allows to identify actin binding proteins in situ. This approach was also applied to reconstruct IFs and is currently used by several groups around the globe.
Structural analysis of integrin mediated cell adhesion and their components
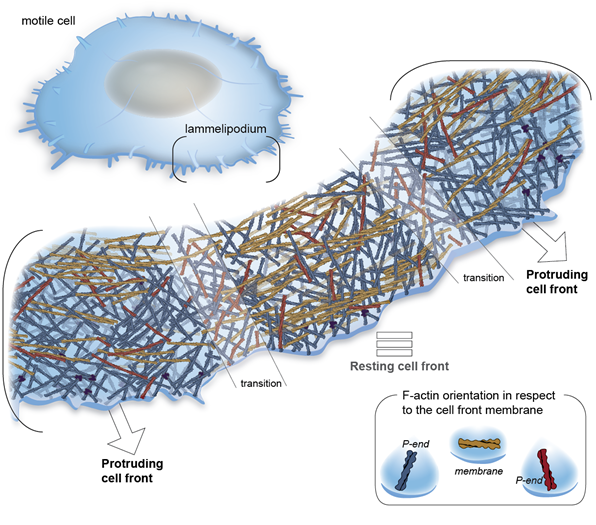
Integrin-based cell adhesions mediate the interactions of the actin cytoskeleton with the extracellular matrix, enabling cell migration, proliferation and differentiation. We have revealed the first cryo-ET map of focal adhesions and revealed the polarity of individual actin filaments at adhesion sites. Using purified proteins, we reveled the function of the full-length vinculin to bundle actin filaments at the onset of adhesion. Moreover, we studied the organization of the cytoskeleton in platelets and reveled the polarity of actin in platelets pseudopodia, as well as the in situ structure of the resting integrin platelets receptor.