Designed Armadillo Repeat Proteins for creating modular, sequence-specific binding to linear epitopes
One of the fundamental prerequisites for almost all biological and biomedical research are specific, well-performing detection agents. To move beyond the unreliable performance of many commercial monoclonal antibodies, we have been working on a completely orthogonal technology. It is aimed at detecting linear epitopes in a completely modular way, amino acid by amino acid, from pre-selected and pre-designed modules, and in a sequence-specific manner.
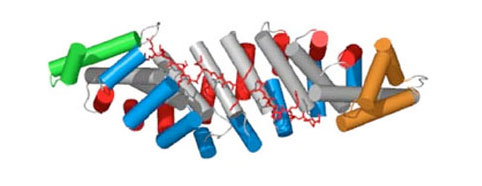
The basis of this technology is synthetic Armadillo Repeat Proteins, which have the ability to bind linear, stretched-out epitopes. A unique feature of this modular binding strategy is that the affinity depends on the length of the peptide and the protein, and thus it can always be adjusted to the «sweet spot» needed for particular applications and experiments.
Using a combination of design, directed evolution and structure determinations by crystallography and NMR, this project brings together all aspects of protein engineering to create the required pockets. These novel pockets have been assembled within Armadillo Repeat Proteins, to confer new sequence-specific binding, and for an ever-increasing number of sequence motifs.