Directed Evolution and Engineering for Stability of GPCRs
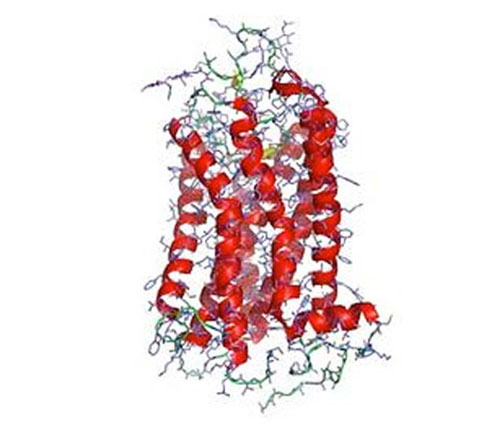
One of the biggest challenges in the study of the structure and biophysics of eukaryotic membrane proteins is that these proteins are extremely difficult to handle: they are usually hard to produce in large amounts, they are very unstable, and they often denature when solubilized in detergent. Our lab has devised several technologies to solve these problems. By expressing mutant libraries of the GPCRs in E. coli and yeast, and by supplying fluorescently labeled agonist or antagonist, those variants can be enriched which express more functional receptor. It has turned out that the variants expressing better in E. coli or yeast also express better in baculovirus-infected insect cells and mammalian cells, and are more stable after solubilization in detergent. Additionally, we have developed a directed evolution technology termed CHESS, in which bacteria are individually wrapped in a polymer, and the inside is completely dissolved by detergent, and thus detergent-stable variants of GPCRs can be selected directly using FACS.
Furthermore, very efficient ways of creating and testing hundreds of GPCR mutants for stability have been devised that have been pivotal in obtaining the recent GPCR structures from our laboratory. Finally, new crystallization chaperones have been created that have ultimately allowed structure determination of a number of recalcitrant receptors. All these efforts have not only made eukaryotic membrane proteins amenable to study, but are helping to elucidate the structural rules of stable membrane proteins.